Introduction. Resistance to carbapenems represents a public health across the world. Screening for carbapenem resistance can be done starting from the antibiogram, but confirmatory methods are usually performed in reference centers. Methods for rapid identification of carbapenemase activity are required to empower laboratories to detect carbapenemases and raise signals for infectious disease physicians, antibiotic stewardship teams and hospital hygiene specialists.
Materials and methods. Eppendorf tubes containing Klebsiella pneumoniae strains with carbapenemase activity (KPC-1, NDM-1, OXA-48) and others with no carbapenemase activity (K. pneumoniae ATCC 700603 and E. coli ATCC 25922), as well as positive and negative controls were incubated at 37 °C with varying concentrations of Meropenem. After incubation, samples were spun down until a pellet formed and the supernatant was used to challenge an exponentially growing E. coli culture. This second culture was followed with nephelometric measurements every half hour. Experiments were carried out in three technical replicates.
The statistical analysis was done with R (version 3.3.3) and graphs generated in RStudio (version 1.0.136) with ggplot2 (version 2.1.0). Mann Whitney U Test with the assumption of independent samples was used to analyse the results.
Results. The doses of Meropenem were chosen after dose ranging experiments and multiple incubation times (from 30 minutes to two hours) were tested in order to evaluate the robustness of the method.
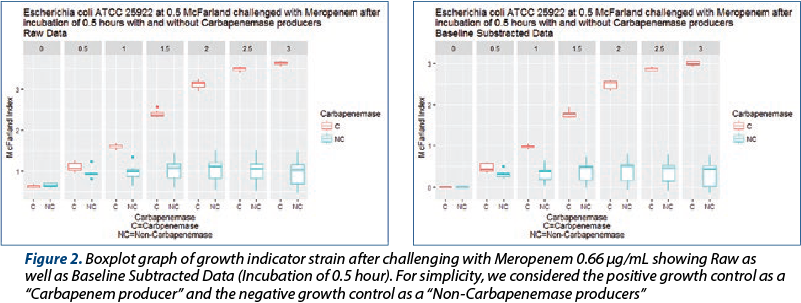
In the cases where the antibiotic was first incubated with strains that had carbapenemase activity, the growth of Escherichia coli ATCC 25 922 was non-hindered, matching that of controls. In the absence of carbapenemase activity and in the case of negative controls, the growth indicator strain was inhibited. Statistically significant differences were identified for 0.3, as well as a 0.5 McFarland index as early as 1.5 hours total work time.
Conclusions. The discovery of carbapenem resistance has direct clinical, as well as epidemiological implications. Among the mechanisms implicated in resistance to carbapenems, the presence of a carbapenemase is one of the most worrying from an epidemiological standpoint. This new tool we describe herein can be performed in all clinical microbiology laboratories, with no extra expense (as opposed to CarbaNP and BlueCarba tests).
Studiu pilot privind o nouă metodă de evaluare a prezenței carbapenemazelor
Carbapenemase Inhibition Challenge Method: a novel way for evaluating the presence of carbapenemases – a pilot study
First published: 11 iulie 2017
Editorial Group: MEDICHUB MEDIA
DOI: 10.26416/Inf.50.2.2017.904
Abstract
Rezumat
Introducere. Rezistența față de carbapeneme este o problemă de sănătate publică pe întregul mapamondul. Screeningul rezistenței față de carbapeneme poate fi făcut începând cu antibiograma, dar metodele de confirmare sunt, de obicei, apanajul centrelor de referință. Metodele rapide de identificare a activității carbapenemazelor sunt necesare pentru a permite laboratoarelor detecția tulpinilor producătoare de carbapenemaze și atragerea atenției medicilor de boli infecțioase, a echipelor de îndrumare antibiotică și a specialiștilor în igienă.
Materiale și metode. Tuburi Eppendorf cu tulpini de Klebsiella pneumoniae producătoare de carbapenemaze (KPC-1, NDM-1, OXA-48) și altele neproducătoare de carbapenemaze (K. pneumoniae ATCC 700603 și E. coli ATCC 25922), precum și controale pozitive și negative au fost incubate la 37 °C cu diferite concentrații de Meropenem. După incubare, tulpinile au fost centrifugate până la formarea unui sediment, iar supernatantul a fost folosit pentru a testa o cultură de E. coli aflată în creștere exponențială. Această a doua cultură a fost folosită pentru măsurătoare nefelometrică la fiecare jumătate de oră. Experimentele au fost executate în trei replicate tehnice. Analiza statistică a fost efectuată folosind R (versiunea 3.3.3), iar graficele au fost generate în RStudio (versiunea 1.0.136) cu ggplot2 (versiunea 2.1.0). Testul Mann Whitney U cu prezumția de independență a eșantioanelor a fost folosit pentru a analiza rezultatele.
Rezultate. Dozele de Meropenem au fost alese în urma unor experimente preliminare; timpul de incubare (variabil de la 30 de minute la două ore) a fost testat pentru a evalua robustețea metodei. În cazurile în care antibioticul a fost întâi incubat cu o tulpină cu activitate de carbapenemază, creșterea tulpinii de Escherichia coli ATCC 25 922 nu a fost inhibată, fiind echivalentă cu cea a controalelor. În absența carbapenemazelor și în cazul controalelor negative, tulpina indicatoare de creștere a fost inhibată. Diferențe semnificative statistic au fost identificate la 1,5 ore din timpul total de lucru, atât când am folosit o cultură indicator de densitate optică inițială de 0,3, cât și una de 0,5 McFarland.
Concluzii. Descoperirea rezistenței față de carbapeneme are atât implicații clinice, cât și epidemiologice. Printre mecanismele implicate în rezistența față de carbapeneme, prezența carbapenemazelor este cea mai îngrijorătoare din punct de vedere epidemiologic. Testul descris în acest articol poate fi efectuat în toate laboratoarele clinice de microbiologie, în absența unor costuri sau dotări suplimentare (spre deosebire de testele CarbaNP și BlueCarba).
Introduction
Carbapenemase-producing Enterobacteriaceae (CPE) are an increasingly growing global threat. Given their plasmidic location, CPE are associated with rapid dissemination in hospital settings and in the community(1-3). These represent a constant pool of resistance genes in the environment(4,5). The frequent association with resistance mechanisms to other classes of antibiotics limits the available therapeutic options(6). This leads to high morbidity and mortality rates.
Rapid identification of carbapenemase production is important clinically, as well as epidemiologically, especially in areas with a high prevalence of CPE. For this purpose, in recent years there have been many attempts of finding a rapid phenotypic test that is both sensitive and specific(7). However, carbapenemases are beta-lactamases that have various levels of expression and the phenotypic aspect may vary considerably(8). The European Committee on Antimicrobial Susceptibility Testing (EUCAST) published in 2013 guidelines regarding the detection of carbapenemase production(9). The phenotypic tests recommended include the combination disk test, carbapenem hydrolysis using MALDI-TOF MS and the CarbaNP test. Meanwhile, other described methods of phenotypic detection of carbapenemase include the Modified Hodge Test, recommended by the CLSI.
In 2015 van der Zwaluw described a new phenotypic method called the Carbapenem Inactivation Method (CIM) as an alternative to the CarbaNP test. CIM is able to detect carbapenemase activity in eight hours(10). This was prospectively evaluated and found to have excellent sensitivity and specificity(11). Yet, the method requires a long incubation period which slows the process of carbapenemase detection and in a setting when information might be used for making decisions pertaining to the patient’s treatment the potential lack of same day results is a disadvantage.
Materials and methods
All bacteria were cultured overnight on Trypticase Soy Agar (TSA) media. Mueller Hinton Broth (MHC) cation adjusted media was incubated at 37 °C under aerobic conditions, with no agitation. Meropenem was preserved as to manufacturer’s indications. Freshly resuspended Meropenem in water was used for each experiment.
Clinical isolates of Klebsiella pneumoniae were obtained from the “Cantacuzino” National Institute for Research. These had characterized carbapenemase activity, harboring: NDM-1, KPC-2, OXA-48.
Reference cultures were used as growth indicator and controls with lack of carbapenemase activity: Klebsiella pneumoniae ATCC 700603 and Escherichia coli ATCC 25922.
A positive growth control was used in the form of Escherichia coli ATCC 25922 challenged with water. Another negative control was done by adding Meropenem in water directly to the Escherichia coli culture.
Modified Minimal Inhibiting Concentration (MMIC) was determined by following the protocol:
- 500 µL of binary dilutions of Meropenem, starting with 64 µg/mL
- Escherichia coli was suspended to a final optical density of 0.5 McFarland units
- 2,500 µL of bacteria was added over each 500 µL of MEM dilution, so as to achieve a final concentration of 64 to 0.125 µg/mL.
The final suspension was incubated for 24 hours, at which time, it was homogenized by vortexing and the McFarland Index was noted.
Carbapenemase Inactivation Challenge Method was performed as follows:
- 2 loopfuls of bacteria grown overnight were resuspended in 1,200 µL of Sterile Water in which Meropenem was dissolved and incubated at 37 °C from 30 minutes up to two hours without shaking.
- After the allotted amount of time, the suspension was pelleted by centrifugation and the supernatant (500 µL) was carefully extracted.
- Before the end of the incubation period, a 2500 µL suspension of Escherichia coli ATCC 25922 at a McFarland Index of 0.5 was prepared. This was then “challenged” with the Meropenem present in the 500 µL of suspension.
Optical densities were monitored every half hour by nephelometry and results were recorded.
All experiments were carried out in triplicate. Results were recorded and imported into R with the addition of RStudio for analysis. Mann-Whitney U test was used to compare the results.
Results
The Modified Minimal Inhibiting Concentration of Escherichia coli ATCC 25922 was 0.5 µg/mL. Initial dose-ranging experiments covered 0.33, 0.5, 0.66 and 1 µg/mL. We used 0.66 µg/mL as minimal Meropenem concentration. This gave consistent discriminatory results. Negative controls (non-carbapenemase producers) showed good discrimination at this level.
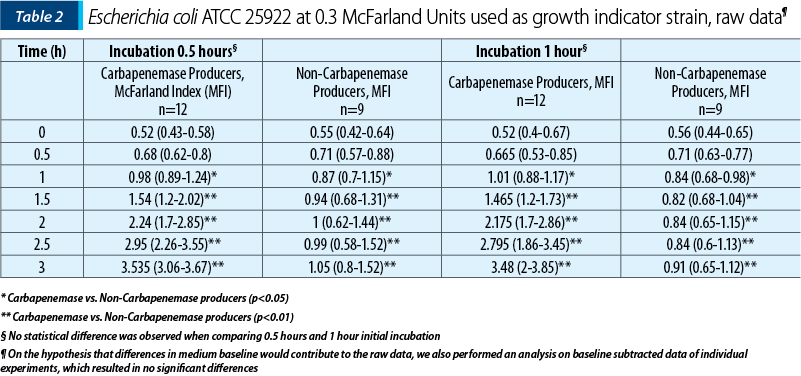
After initial experiments in order to find the time needed for incubation, we experimented for 30 minutes and one hour long incubation. After incubation and pelleting, 500 µL of supernatant was used to challenge a freshly suspended culture of 0.5 McFarland of Escherichia coli ATCC 25922. Results of growth monitoring are presented in graphical and tabular form.
Due to homogeneity of results, we utilized solely 30 minute and 1 hour incubation afterwards (see Figures 1 and 2). For the purposes of display, a jitter plot was used due to the superposition of numerous experiments.
While, after one hour of growth, statistically significant differences could be determined between carbapenem producers and non-producers (Table 1 and Table 2), we believe that the small difference is not operational – that is, could not be detected and interpreted correctly in light of the small difference (approximately 0.1 McFarland units). We thus suggest interpretation to be done at 1.5 hours of incubation, where the median difference was more than one McFarland unit and the minimal difference was 0.5 McFarland units.
Discussions
We present herein a pilot study of a novel protocol used to easily and rapidly detect carbapenemase producer strains. Methodologically, this study builds on two different concepts.
First concept is the use of a low carbapenem dose as challenger. The Enterobacteriaceae strains most difficult to identify are the OXA-48 and OXA-48-like producers which have a relatively weak lytic effect on carbapenems(12). Then, there are the NDM strains, for which the addition of ZnSO4 has been proposed as a technical solution(13). The slow lysis capacity may give false negative results through the incapacity to generate the large amounts of acidic compound required to produce a color shift in the working solution (be it either Phenol Red or Methylene Blue)(14). Yet, it has been shown that the same enzyme has different phenotypic impact dependent on the level of expression and the permeability of the membrane(15).
The second concept is the use of an exponentially growing indicator strain available to all laboratories (we can use Escherichia coli ATCC 25922 or another isolate, which can be customized by the particular laboratory). Further work is needed to optimize our understanding of Escherichia coli growth in the presence of carbapenemases.
Interestingly, we found no need for ZnSO4 in our small batch of experiments. It’s possible that this may come into play with higher doses of carbapenems.
The limitations of this study have to take into account the lack of other types of negative controls. We used a well characterized Klebsiella pneumoniae strain which lacks a carbapenemase but that has an ESBL (Extended Spectrum Beta Lactamase). We rejected one experiment in which the negative controls had unusually high growth levels. We believe that this may be related to the instability of Meropenem (and all other carbapenems) when resuspended(16). On the other hand, results can be easily checked by the use of positive and negative controls. We used analytical grade carbapenem, but, as showed, injectable carbapenem works well. Further investigation is required to confirm this hypothesis(17).
Conclusions
Rapid detection of CPE is extremely important in choosing the right empirical therapeutic option and in preventing outbreaks. Our proposed test, the Carbapenem Inactivation Challenge Method (CICM), has the perspective of being a cheaper and at least equally rapid test to those commercially available. Further studies are necessary to assess the sensitivity and specificity of the test compared with what is already on the market and to consider multiple bacterial strains with different types of carbapenemases and other factors that may contribute to resistance (like porin mutations).
Conflict of interests: The authors declare no conflict of interests.
Bibliografie
- van Duin D, Doi Y. The global epidemiology of carbapenemase-producing Enterobacteriaceae. Virulence. 2016;11:1-10.
- Tang H-J, Hsieh C-F, Chang P-C, et al. Clinical Significance of Community- and Healthcare-Acquired Carbapenem-Resistant Enterobacteriaceae Isolates. PLoS ONE. 2016;21:11(3):e0151897.
- Grundmann H, Glasner C, Albiger B, et al. Occurrence of carbapenemase-producing Klebsiella pneumoniae and Escherichia coli in the European survey of carbapenemase-producing Enterobacteriaceae (EuSCAPE): a prospective, multinational study. Lancet Infect Dis. 2017;17(2):153-63.
- Friedman ND, Carmeli Y, Walton AL, Schwaber MJ. Carbapenem-Resistant Enterobacteriaceae: A Strategic Roadmap for Infection Control. Infect Control Hosp Epidemiol. 2017;15:1-15.
- Nordmann P, Naas T, Poirel L. Global spread of Carbapenemase-producing Enterobacteriaceae. Emerging Infect Dis. 2011;17(10):1791-8.
- Falagas ME, Tansarli GS, Karageorgopoulos DE, Vardakas KZ. Deaths attributable to carbapenem-resistant Enterobacteriaceae infections. Emerging Infect Dis. 2014;20(7):1170-5.
- Kost K, Yi J, Rogers B, Jerris R. Comparison of clinical methods for detecting carbapenem-resistant Enterobacteriaceae. Practical Lab Med. 2017;8:18-25.
- Queenan AM, Bush K. Carbapenemases: the versatile beta-lactamases. Clin Microbiol Rev. 2007;20(3):440-58
- EUCAST. EUCAST guidelines for detection of resistance mechanisms and specific resistances of clinical and/or epidemiological importance. EUCAST guidelines for detection of resistance mechanisms and specific resistances of clinical and/or epidemiological importance. 2013 [cited 2017 Mar 30]. Available from: http://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Resistance_mechanisms/EUCAST_detection_of_resistance_mechanisms_v1.0_20131211.pdf
- van der Zwaluw K, de Haan A, Pluister GN, et al. The carbapenem inactivation method (CIM), a simple and low-cost alternative for the Carba NP test to assess phenotypic carbapenemase activity in gram-negative rods. PLoS ONE. 2015;23,10(3):e0123690.
- Gauthier L, Bonnin RA, Dortet L, Naas T. Retrospective and prospective evaluation of the Carbapenem inactivation method for the detection of carbapenemase-producing Enterobacteriaceae. PLoS ONE. 20173;12(2):e0170769.
- Bakthavatchalam YD, Anandan S, Veeraraghavan B. Laboratory Detection and Clinical Implication of Oxacillinase-48 like Carbapenemase: The Hidden Threat. J Glob Infect Dis. 2016;8(1):41-50.
- Girlich D, Poirel L, Nordmann P. Value of the modified Hodge test for detection of emerging carbapenemases in Enterobacteriaceae. J Clin Microbiol. 2012;50(2):477-9.
- Wang Y, Lo W-U, Yau C-Y, et al. Impact of intraobserver and interobserver variation on performance of the CLSI Carba NP assay for carbapenemase detection in Enterobacteriaceae. J Glob Antimicrob Resist. 2017;16;9:19-20.
- Antunes NT, Lamoureaux TL, Toth M, et al. Class D β-lactamases: are they all carbapenemases? Antimicrob Agents Chemother. 2014;58(4):2119-25.
- Tomasello C, Leggieri A, Cavalli R, Di Perri G, D’Avolio A. In vitro stability evaluation of different pharmaceutical products containing meropenem. Hosp Pharm. 2015;50(4):296-303.
- Pires J, Novais A, Peixe L. Blue-carba, an easy biochemical test for detection of diverse carbapenemase producers directly from bacterial cultures. J Clin Microbiol. 2013;51(12):4281-3.